bwbioinfo · GitHub
Last updated:2026-03-13 12:43
gganimate-example
Animation
old_wd <- setwd(dirname(knitr::current_input()))
on.exit(setwd(old_wd), add = TRUE)
# Load required libraries
invisible(lapply(required_pkgs, library, character.only = TRUE))
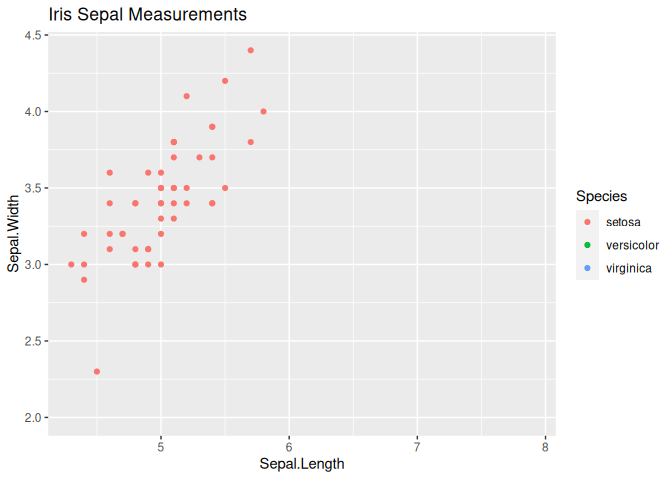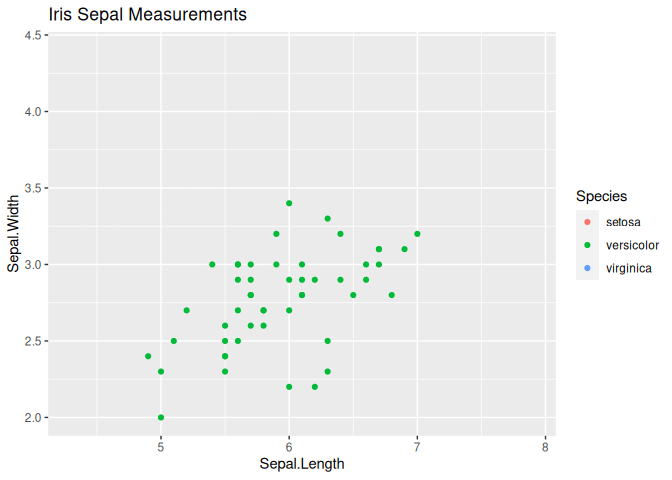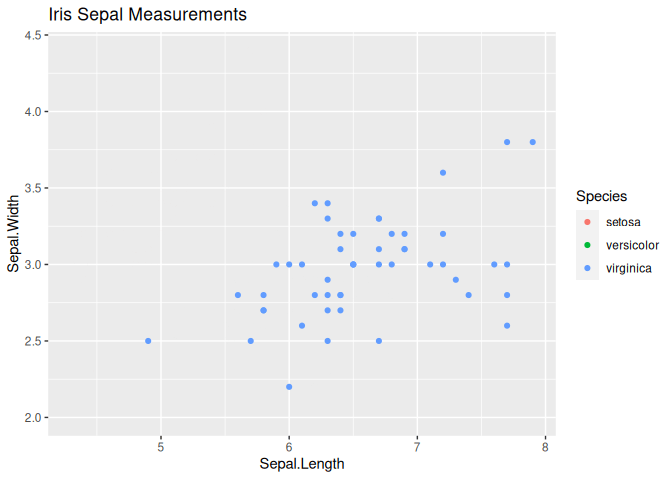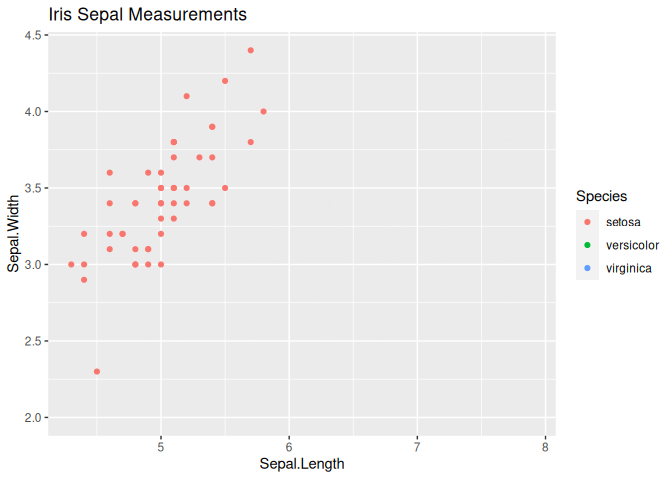
# Create a static plot
p <-
ggplot(
iris,
aes(
x = Sepal.Length,
y = Sepal.Width,
color = Species)
) +
geom_point() +
labs(
title = 'Iris Sepal Measurements'
)
# Animate the plot by transitioning through Species
anim <-
p + transition_states(
Species,
transition_length = 2,
state_length = 1
) +
enter_fade() +
exit_fade()
# Render
animate(anim, nframes = 100, fps = 10)
Session Info
sessionInfo()
## R version 4.5.2 (2025-10-31)
## Platform: x86_64-pc-linux-gnu
## Running under: Linux Mint 22.2
##
## Matrix products: default
## BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.12.0
## LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.12.0 LAPACK version 3.12.0
##
## locale:
## [1] LC_CTYPE=en_CA.UTF-8 LC_NUMERIC=C
## [3] LC_TIME=en_CA.UTF-8 LC_COLLATE=en_CA.UTF-8
## [5] LC_MONETARY=en_CA.UTF-8 LC_MESSAGES=en_CA.UTF-8
## [7] LC_PAPER=en_CA.UTF-8 LC_NAME=C
## [9] LC_ADDRESS=C LC_TELEPHONE=C
## [11] LC_MEASUREMENT=en_CA.UTF-8 LC_IDENTIFICATION=C
##
## time zone: America/Toronto
## tzcode source: system (glibc)
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## loaded via a namespace (and not attached):
## [1] vctrs_0.7.1 cli_3.6.5 knitr_1.51 rlang_1.1.7
## [5] xfun_0.56 otel_0.2.0 generics_0.1.4 S7_0.2.1
## [9] jsonlite_2.0.0 glue_1.8.0 htmltools_0.5.9 sass_0.4.10
## [13] scales_1.4.0 rmarkdown_2.30 grid_4.5.2 tibble_3.3.1
## [17] evaluate_1.0.5 jquerylib_0.1.4 fastmap_1.2.0 yaml_2.3.12
## [21] lifecycle_1.0.5 bookdown_0.46 compiler_4.5.2 dplyr_1.2.0
## [25] RColorBrewer_1.1-3 pkgconfig_2.0.3 blogdown_1.23 farver_2.1.2
## [29] digest_0.6.39 R6_2.6.1 tidyselect_1.2.1 pillar_1.11.1
## [33] magrittr_2.0.4 bslib_0.10.0 tools_4.5.2 gtable_0.3.6
## [37] ggplot2_4.0.2 cachem_1.1.0
