bwbioinfo · GitHub
Last updated:2026-03-13 12:43
Cumulative Reads Over Time
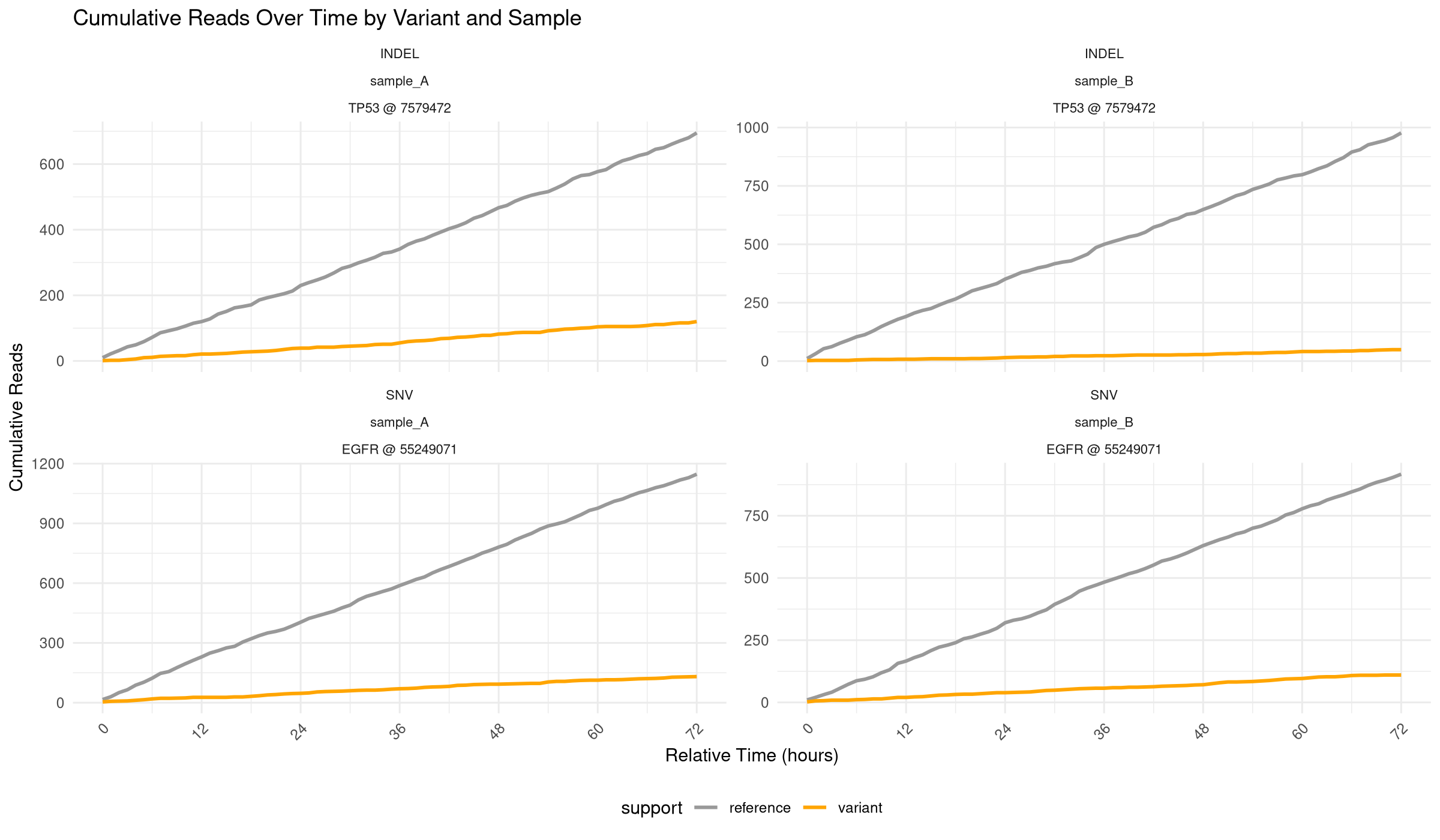
This script isolates the cumulative read-trajectory panel, matching the 72-hour variant/reference trend view from the full analysis workflow. This code reproduces the base graphic reported in the article Single-workflow Nanopore whole genome sequencing with adaptive sampling for accelerated and comprehensive pediatric cancer profiling.
Data setup
library(dplyr)
library(tidyr)
library(ggplot2)
set.seed(42)
time_hours <- seq(0, 72, by = 1)
samples <- c("sample_A", "sample_B")
variants <- c("EGFR @ 55249071", "TP53 @ 7579472")
support_types <- c("variant", "reference")
agg_cum <- tidyr::crossing(
sample_name = samples,
var_region = variants,
support = support_types,
rel_time_hours = time_hours
) %>%
group_by(sample_name, var_region, support) %>%
mutate(
rate = ifelse(
support == "variant",
runif(1, 0.7, 2.0),
runif(1, 8, 16)
),
incremental = rpois(n(), lambda = rate),
cumulative_reads = cumsum(incremental),
variant_type = ifelse(
grepl("EGFR", var_region),
"SNV",
"INDEL"
)
) %>%
ungroup()
Isolated plot
p_main <- ggplot(
agg_cum,
aes(
x = rel_time_hours,
y = cumulative_reads,
color = support
)
) +
geom_line(linewidth = 1) +
scale_color_manual(
values = c(
"variant" = "orange",
"reference" = "gray60"
),
labels = c(
"reference" = "Reference Reads",
"variant" = "Variant Reads"
)
) +
scale_x_continuous(
breaks = seq(0, 72, by = 12),
minor_breaks = seq(0, 72, by = 6)
) +
facet_wrap(
variant_type ~ sample_name + var_region,
scales = "free_y",
drop = TRUE
) +
labs(
title = paste(
"Cumulative Reads Over Time",
"by Variant and Sample"
),
subtitle = paste(
"Orange lines show variant-supporting reads,",
"gray lines show reference reads"
),
x = "Relative Time (hours from sequencing start)",
y = "Cumulative Reads"
) +
theme_minimal() +
theme(
legend.position = "bottom",
strip.text = element_text(size = 8),
axis.text.x = element_text(angle = 45, hjust = 1)
)
p_main
Session Info
## R version 4.5.2 (2025-10-31)
## Platform: x86_64-pc-linux-gnu
## Running under: Linux Mint 22.2
##
## Matrix products: default
## BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.12.0
## LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.12.0 LAPACK version 3.12.0
##
## locale:
## [1] LC_CTYPE=en_CA.UTF-8 LC_NUMERIC=C
## [3] LC_TIME=en_CA.UTF-8 LC_COLLATE=en_CA.UTF-8
## [5] LC_MONETARY=en_CA.UTF-8 LC_MESSAGES=en_CA.UTF-8
## [7] LC_PAPER=en_CA.UTF-8 LC_NAME=C
## [9] LC_ADDRESS=C LC_TELEPHONE=C
## [11] LC_MEASUREMENT=en_CA.UTF-8 LC_IDENTIFICATION=C
##
## time zone: America/Toronto
## tzcode source: system (glibc)
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## loaded via a namespace (and not attached):
## [1] digest_0.6.39 R6_2.6.1 bookdown_0.46 fastmap_1.2.0
## [5] xfun_0.56 blogdown_1.23 cachem_1.1.0 knitr_1.51
## [9] htmltools_0.5.9 rmarkdown_2.30 lifecycle_1.0.5 cli_3.6.5
## [13] sass_0.4.10 jquerylib_0.1.4 compiler_4.5.2 tools_4.5.2
## [17] evaluate_1.0.5 bslib_0.10.0 yaml_2.3.12 otel_0.2.0
## [21] jsonlite_2.0.0 rlang_1.1.7
ggplot2 nanopore cumulative reads variant
392 Words
2026-03-07 19:00 (Last updated: 2026-03-13 12:43)
