bwbioinfo · GitHub
Last updated:2026-03-13 12:43
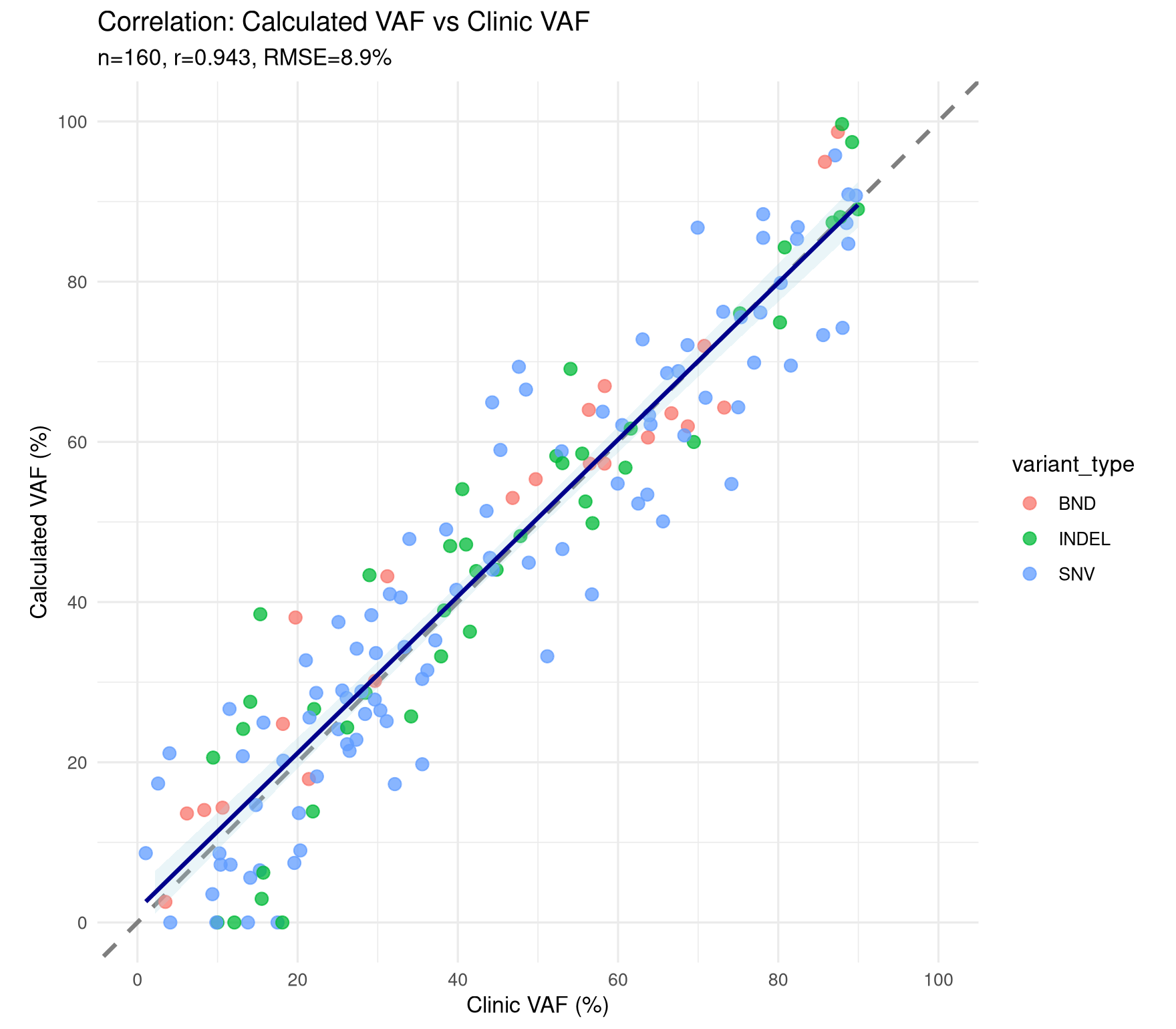
VAF Correlation Scatter
This script isolates the clinic-vs-calculated VAF correlation panel, matching the scatter/regression view from the full analysis workflow. This code reproduces the base graphic reported in the article Single-workflow Nanopore whole genome sequencing with adaptive sampling for accelerated and comprehensive pediatric cancer profiling.
Data setup
library(dplyr)
library(ggplot2)
set.seed(123)
vaf_correlation_data <- tibble::tibble(
sample_name = sample(
paste0("sample_", LETTERS[1:8]),
160,
replace = TRUE
),
variant_type = sample(
c("SNV", "INDEL", "BND"),
160,
replace = TRUE,
prob = c(0.6, 0.25, 0.15)
),
clinic_vaf = runif(160, min = 1, max = 90)
) %>%
mutate(
vaf = pmin(
100,
pmax(0, clinic_vaf + rnorm(n(), mean = 0, sd = 9))
),
vaf_diff = vaf - clinic_vaf
)
correlation_stats <- vaf_correlation_data %>%
summarise(
n = n(),
correlation = cor(vaf, clinic_vaf),
rmse = sqrt(mean(vaf_diff^2))
)
Isolated plot
p_vaf_correlation <- vaf_correlation_data %>%
ggplot(aes(x = clinic_vaf, y = vaf)) +
geom_abline(
intercept = 0,
slope = 1,
linetype = "dashed",
color = "gray50",
size = 1
) +
geom_point(
aes(color = variant_type),
size = 2.6,
alpha = 0.75
) +
geom_smooth(
method = "lm",
se = TRUE,
color = "darkblue",
fill = "lightblue",
alpha = 0.25
) +
scale_x_continuous(
limits = c(0, 100),
breaks = seq(0, 100, 20)
) +
scale_y_continuous(
limits = c(0, 100),
breaks = seq(0, 100, 20)
) +
coord_fixed() +
labs(
title = "Correlation: Calculated VAF vs Clinic VAF",
subtitle = sprintf(
"n=%d, r=%.3f, RMSE=%.1f%%",
correlation_stats$n,
correlation_stats$correlation,
correlation_stats$rmse
),
x = "Clinic VAF (%)",
y = "Calculated VAF (%)"
) +
theme_minimal()
p_vaf_correlation
Session Info
## R version 4.5.2 (2025-10-31)
## Platform: x86_64-pc-linux-gnu
## Running under: Linux Mint 22.2
##
## Matrix products: default
## BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.12.0
## LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.12.0 LAPACK version 3.12.0
##
## locale:
## [1] LC_CTYPE=en_CA.UTF-8 LC_NUMERIC=C
## [3] LC_TIME=en_CA.UTF-8 LC_COLLATE=en_CA.UTF-8
## [5] LC_MONETARY=en_CA.UTF-8 LC_MESSAGES=en_CA.UTF-8
## [7] LC_PAPER=en_CA.UTF-8 LC_NAME=C
## [9] LC_ADDRESS=C LC_TELEPHONE=C
## [11] LC_MEASUREMENT=en_CA.UTF-8 LC_IDENTIFICATION=C
##
## time zone: America/Toronto
## tzcode source: system (glibc)
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## loaded via a namespace (and not attached):
## [1] digest_0.6.39 R6_2.6.1 bookdown_0.46 fastmap_1.2.0
## [5] xfun_0.56 blogdown_1.23 cachem_1.1.0 knitr_1.51
## [9] htmltools_0.5.9 rmarkdown_2.30 lifecycle_1.0.5 cli_3.6.5
## [13] sass_0.4.10 jquerylib_0.1.4 compiler_4.5.2 tools_4.5.2
## [17] evaluate_1.0.5 bslib_0.10.0 yaml_2.3.12 otel_0.2.0
## [21] jsonlite_2.0.0 rlang_1.1.7
